Pham, K.M. and P.A. Beal. “Structure-Guided Optimization of siRNA and Anti-miRNA Properties.” In: Sugimoto, N. (eds) Handbook of Chemical Modifications of Nucleic Acids. Springer, Singapore 2023. DOI:10.1007/978-981-16-1313-5_41-1
Pham, K.M.; Suter, S.R.; Lu, S.S.; Beal, P.A. Ester modification at the 3′ end of anti-microRNA oligonucleotides increases potency of microRNA inhibition. Bioorganic & Medicinal Chemistry 2020. DOI:10.1016/j.bmc.2020.115894
Hu, T.; Suter, S. R.; Mumbleau, M. M.; Beal, P. A., TLR8 activation and inhibition by guanosine analogs in RNA: Importance of functional groups and chain length. Bioorganic & Medicinal Chemistry 2017.
DOI:10.1016/j.bmc.2017.11.020
Suter, S. R.; Ball-Jones, A.; Mumbleau, M. M.; Valenzuela, R.; Ibarra-Soza, J.; Owens, H.; Fisher, A. J.; Beal, P. A., “Controlling miRNA-like off-target effects of an siRNA with nucleobase modifications.” Organic & Biomolecular Chemistry 2017.
DOI:10.1039/C7OB02654D
Valenzuela, R. A. P., et al. (2016). “Guide Strand 3′-End Modifications Regulate siRNA Specificity.” ChemBioChem 17(24): 2340-2345. DOI: 10.1002/cbic.201600453
Suter, S. R., et al. (2016). “Structure-Guided Control of siRNA Off-Target Effects.” Journal of the American Chemical Society 138(28): 8667-8669. DOI: 10.1021/jacs.6b06137
Zheng, Y. and P. A. Beal (2016). “Synthesis and evaluation of an alkyne-modified ATP analog for enzymatic incorporation into RNA.” Bioorganic & Medicinal Chemistry Letters 26(7): 1799-1802. DOI: 10.1016/j.bmcl.2016.02.038
Valenzuela, R. A. P., et al. (2015). “Base Modification Strategies to Modulate Immune Stimulation by an siRNA.” ChemBioChem 16(2): 262-267. DOI: 10.1002/cbic.201402551
Kiet Tran, Michelle R. Arkin, and Peter A. Beal, “Tethering in RNA: An RNA-Binding Fragment Discovery Tool”, Molecules, 2015,20, 4148-4161. DOI:10.3390/molecules20034148
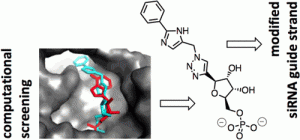
Onizuka, K., et al. (2013). “Short Interfering RNA Guide Strand Modifiers from Computational Screening.” Journal of the American Chemical Society 135(45): 17069-17077. DOI: 10.1021/ja4079754
-Highlighted in the Chemical & Engineering News, News of the Week, Nov. 18 2013 issue, “Gene Silencing by Design”
-Highlighted in ACS Chemical Biology Spotlight, December 20, 2013 issue, “Running Interference with Synthetic SiRNAs”
-Highlighted in SciBX:Science-Business exchange (Nature Publishing), December 19, 2013: “Modifications to the siRNA guide strand that improve therapeutic potency”
Harrison, J. G., et al. (2013). “Computational Approaches to Predicting the Impact of Novel Bases on RNA Structure and Stability.” ACS Chemical Biology 8(11): 2354-2359. DOI:10.1021/cb4006062
Ghanty, U., et al. (2012). “Promiscuous 8-Alkoxyadenosines in the Guide Strand of an SiRNA: Modulation of Silencing Efficacy and Off-Pathway Protein Binding.” Journal of the American Chemical Society 134(42): 17643-17652. DOI:10.1021/ja307102g
Ibarra-Soza, J. M., et al. (2012). “7-Substituted 8-aza-7-deazaadenosines for modification of the siRNA major groove.” Organic & Biomolecular Chemistry 10(32): 6491-6497. DOI:10.1039/C2OB25647A
“8-Oxoguanosine switches modulate the activity of alkylated siRNAs by controlling steric effects in the major vs. minor grooves” Arunkumar Kannan, Erik Fostvedt, Peter A. Beal and Cynthia J. Burrows J. Am. Chem. Soc 2011, 133, 6343-6351.
“Covalent stabilization of a small molecule-RNA complex” Hayden Peacock, Radhika Bachu, and Peter A. Beal Bioorg. Med. Chem. Lett. 2011, 21, 5002-5005.
“Nucleobase and ribose modifications control immunostimulation of a microRNA 122-mimetic siRNA” Hayden Peacock, Raymond V. Fucini, Prasanna Jayalath, José M. Ibarra-Soza, Henry J. Haringsma, W. Michael Flanagan, Aarron Willingham and Peter A. Beal J. Am. Chem. Soc. 2011, 133, 24, 9200-9203.
-Highlighted in ACS Chemical Biology Spotlight, July 15, 2011 issue, “Minimizing MicroRNA Immunostimulation”
-Selected as a Faculty of 1000 highlighted paper
-Highlighted in Nov. 1, 2011 issue of Genetic Engineering and Biotechnology News
“Chemical Modification of siRNA Bases to Probe and Enhance RNA Interference” Hayden Peacock, Arunkumar Kannan, Peter A. Beal and Cynthia J. Burrows J. Org. Chem. 2011, 76, 18, 7295-7300.
“Minor-groove-modulating adenosine replacements control protein binding and RNAi activity in siRNAs” Hayden Peacock, Erik Fostvedt and Peter A. Beal ACS Chem. Biol. 2010, 5, 12, 1115-1124.
“N2-Modified 2-aminopurine ribonucleotides as minor groove-modulating adenosine replacements in duplex RNA” Hayden Peacock, Olena Maydanovych and Peter A. Beal Org. Lett. 2010, 12, 5, 1044-1047.
